It is clear, however, that segmenting one clownfish with particular lighting and background may not necessarily generalize well to segmenting all clownfish. Overall, this simple segmentation method has successfully located the majority of Nemo’s relatives. The shadowed bottom half of Nemo’s nephew is completely excluded, but bits of the purple anemone in the background look awfully like Nemo’s blue tinged stripes... The foreground clownfish has orange shades darker than our range. To convert an image from RGB to HSV, you can use cvtColor():įor i in range ( 1, 6 ): plt. The third axis, saturation, defines the shades of hue from least saturated, at the vertical axis, to most saturated furthest away from the center: Image: Wikipedia Values go from dark (0 at the bottom) to light at the top. The colors, or hues, are modeled as an angular dimension rotating around a central, vertical axis, which represents the value channel. We saw Nemo in RGB space, so now let’s view him in HSV space and compare.Īs mentioned briefly above, HSV stands for Hue, Saturation, and Value (or brightness), and is a cylindrical color space. Since parts of Nemo stretch over the whole plot, segmenting Nemo out in RGB space based on ranges of RGB values would not be easy. Here is the colored scatter plot for the Nemo image in RGB:įrom this plot, you can see that the orange parts of the image span across almost the entire range of red, green, and blue values. flatten (), facecolors = pixel_colors, marker = "." ) > axis.

Note that while the current version of OpenCV is 3.x, the name of the package to import is still cv2: This article will assume you have Python 3.x installed on your system. Color Spaces and Reading Images in OpenCVįirst, you will need to set up your environment. If you are not familiar with NumPy or Matplotlib, you can read about them in the official NumPy guide and Brad Solomon’s excellent article on Matplotlib. Slightly different versions won’t make a significant difference in terms of following along and grasping the concepts. This articles uses OpenCV 3.2.0, NumPy 1.12.1, and Matplotlib 2.0.2. The key Python packages you’ll need to follow along are NumPy, the foremost package for scientific computing in Python, Matplotlib, a plotting library, and of course OpenCV. Let’s see how well we can find Nemo in an image. Clownfish are easily identifiable by their bright orange color, so they’re a good candidate for segmentation.
#Python colors download#
To demonstrate the color space segmentation technique, we’ve provided a small dataset of images of clownfish in the Real Python materials repository here for you to download and play with. Remove ads Simple Segmentation Using Color Spaces Now that we understand the concept of color spaces, we can go on to use them in OpenCV. Color spaces are fully able to represent all the colors we are able to distinguish between. Color spaces, however, represent color through discrete structures (a fixed number of whole number integer values), which is acceptable since the human eye and perception are also limited. In reality, color is a continuous phenomenon, meaning that there are an infinite number of colors.
#Python colors software#
These color spaces are frequently used in color selection tools in software and for web design. HSV and HSL are descriptions of hue, saturation, and brightness/luminance, which are particularly useful for identifying contrast in images. They can be analyzed in HED space, a representation of the saturations of the stain types-hematoxylin, eosin, and DAB-applied to the original tissue.
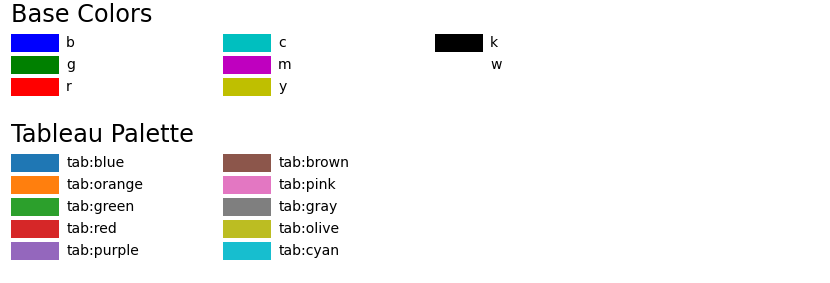
In certain types of medical fields, glass slides mounted with stained tissue samples are scanned and saved as images. Our printers contain ink canisters of cyan, magenta, yellow, and black.
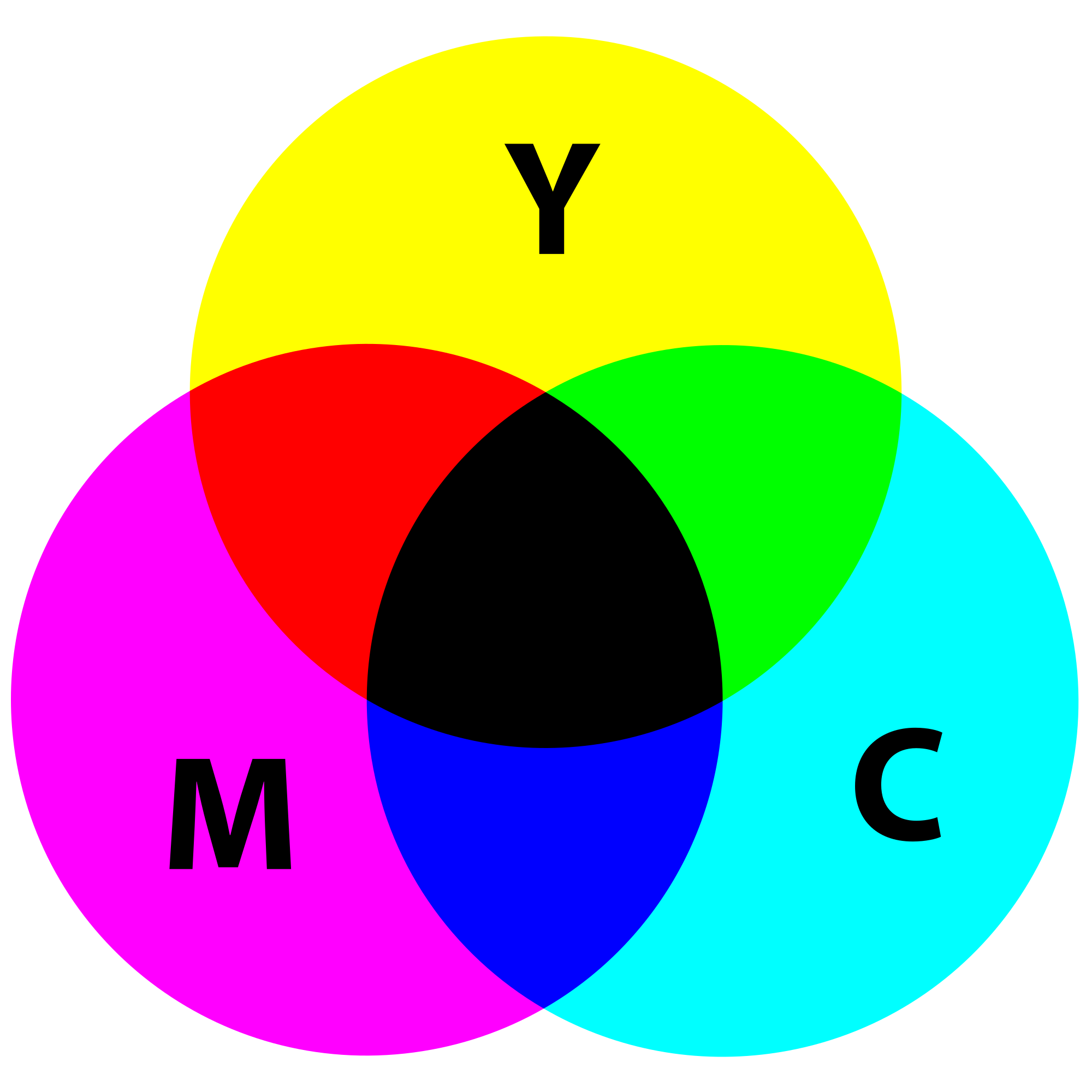
While the 0 tuple in RGB is black, in CMYK the 0 tuple is white. In the printing world, CMYK is useful because it describes the color combinations required to produce a color from a white background. There are so many color spaces because different color spaces are useful for different purposes. RGB is one of the five major color space models, each of which has many offshoots.
